Analyzing the cell cycle is crucial for understanding proliferation and disease. While high-throughput methods sacrifice spatial information, Cytely enables rapid classification of cell cycle stages using only a nuclear stain and morphological profiling, preserving spatial context and enabling reproducible quantification.
Downloadable Files
Cell Cycle Analysis in HeLa Cells
This application note demonstrates how Cytely classifies cell cycle stages using morphological features derived from a standard nuclear stain in adherent HeLa cells. The approach combines automated segmentation with interactive gating to isolate a pure mitotic population while preserving spatial context.
Key features demonstrated in this note:
- Visualizing morphological heterogeneity across a cell population
- Identifying cell cycle phases (G0/G1, S/G2, M) using nuclear morphology
- Sequential gating to isolate and quantify mitotic cells with high purity
- Complex cell cycle analysis without specialized dyes; supports retrospective analysis
Experimental Method
HeLa cells were cultured and stained with a standard nuclear dye. Imaging was performed on adherent cells to preserve morphology. Cytely processed images using automated object detection (nuclei) and measurement of morphological parameters (e.g., area, circularity, major/minor axes, intensity).
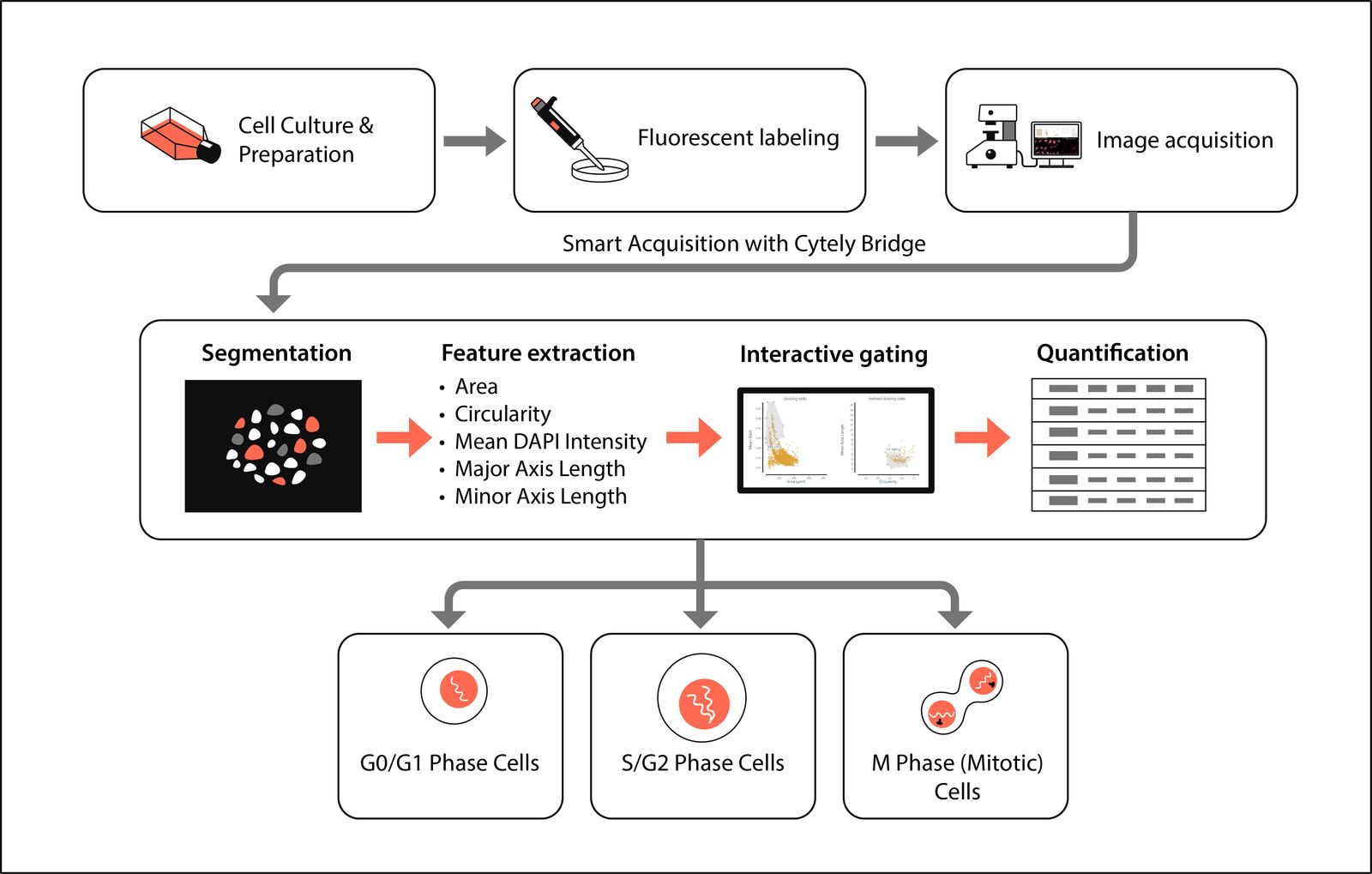
Data Analysis in Cytely
- Upload: Images were added via Cytely’s web interface.
- Segmentation: Automated detection of nuclei and measurement of morphology and intensities.
- Sequential scatter plots: Size- and intensity-derived proxies separated G0/G1, S/G2, and mitotic populations.
- Mitotic isolation: A final gate on Major vs. Minor Axis Length isolated elongated, dividing cells.
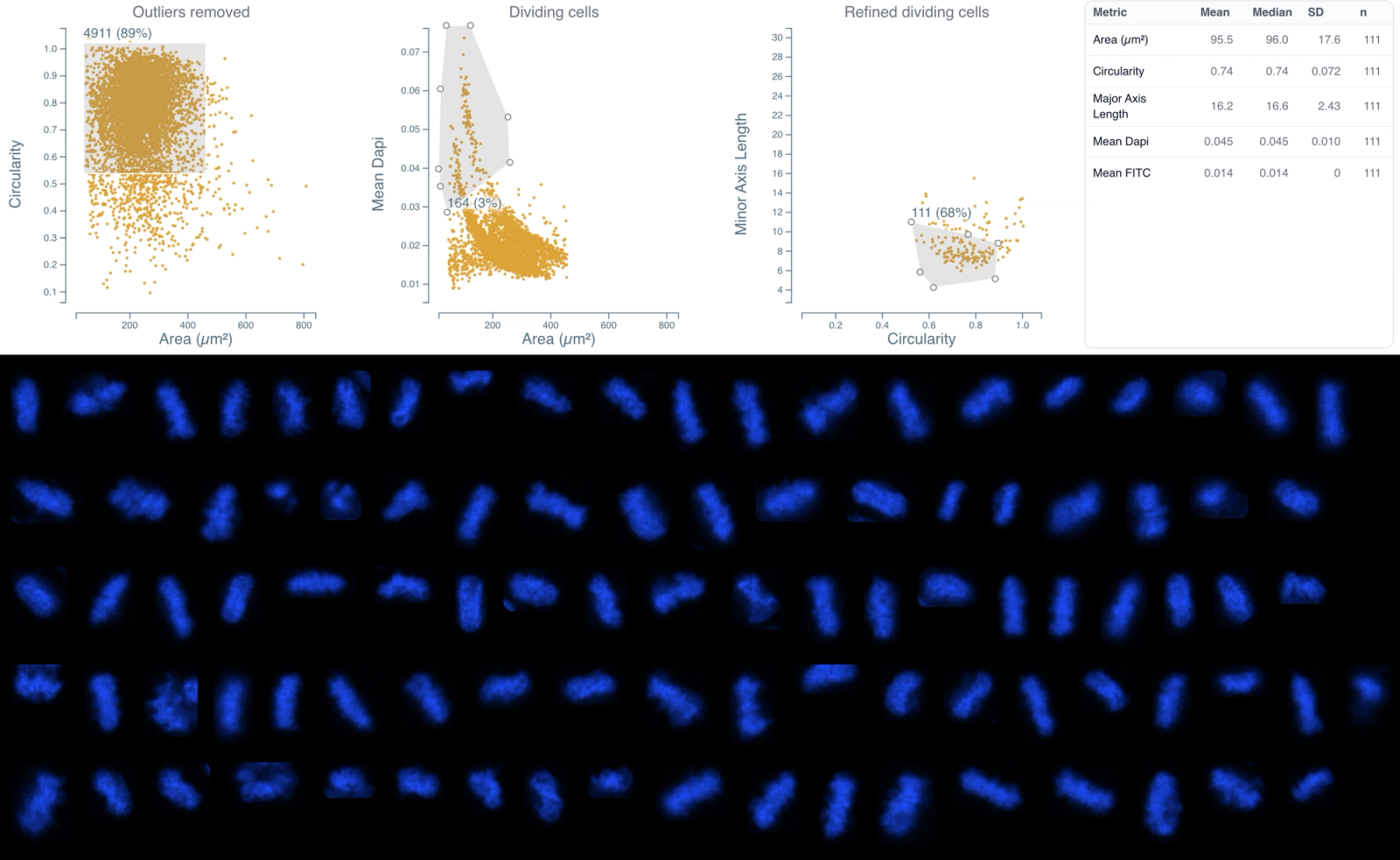
Results
Three major populations were classified by morphology with preserved spatial context. Representative crops of gated cells confirm phenotype by visual inspection. Quantification across the dataset:
- G0/G1: 65%
- S/G2: 23%
- Mitosis: 3%
- Other: 9%

Conclusions
Cytely transforms cell cycle analysis into a fast, intuitive workflow. Automated segmentation combined with multi-parameter gating enables precise quantification of cell cycle stages using standard nuclear staining. The workflow preserves spatial context and provides immediate visual confirmation, yielding reproducible, publication-ready results.
- Eliminates complex multi-dye protocols
- Enables retrospective analysis of existing datasets
- Preserves spatial context that flow cytometry loses
- Immediate visual validation of gated populations
